
Jason J. Corso
| Snippets by Topic | |
| * | Active Clustering |
| * | Activity Recognition |
| * | Medical Imaging |
| * | Metric Learning |
| * | Semantic Segmentation |
| * | Video Segmentation |
| * | Video Understanding |
| Selected Project Pages | |
| * | Action Bank |
| * | LIBSVX: Supervoxel Library and Evaluation |
| * | Brain Tumor Segmentation |
| * | CAREER: Generalized Image Understanding |
| * | Summer of Code 2010: The Visual Noun |
| * | ACE: Active Clustering |
| * | ISTARE: Intelligent Spatiotemporal Activity Reasoning Engine |
| * | GBS: Guidance by Semantics |
| * | Semantic Video Summarization |
| Data Sets | |
| * | YouCook |
| * | Chen Xiph.org |
| * | UB/College Park Building Facades |
| Other Information | |
| * | Code/Data Downloads |
| * | List of Grants |
Joint Segmentation and Classification of Brain Tumors in 3D MRI
People: Jason Corso, Dheerendra Prasad (RPCI)
Past Members Alan Yuille (UCLA), Usha Sinha (UCLA), Shishir Dube (UCLA), Timothy Burns There is thus currently a great deal of interest in the establishment of imaging as a valid biomarker in the assessment of the response of cancer to a variety of treatments [Bradley et al., 2002, Tuma, 2006]. Of the possible measure- ments that can be made on images to assess the change in tumor burden, overall tumor volume is the key indicator [Bradley et al., 2002, Liu et al., 2005]. However, rather than using the full tumor volume, the current international standard measurements rely on basic quantitative surrogates such as the two dimensional World Health Organiza- tion (WHO) [World Health Organization, 1979] or Response Assessment Criteria for Solid Tumors (RECIST) one dimensional criterion [Therasse et al., 2000] used to measure patient response. Specifically, tumors are measured in one or two dimensions at their baseline�the area of maximal size, which is selected manually by the physician. These current quantitative techniques, although offering numerical representations of the state of the patient, are gross approximations to the true underlying tumor and are computed manually, hence time consuming and often not reproducible. They are therefore considered by many to be inadequate to allow imaging to serve as a biomarker on which to base treatment [Patz Jr. et al., 2000, Tuma, 2006]; they have generally failed to provide repeatable and statistically meaningful prognostic parameters of outcome prediction in both the clinic and the research lab. Hence, advanced image segmentation and recognition techniques are needed to improved the measurement science of cancer imaging and tumor staging.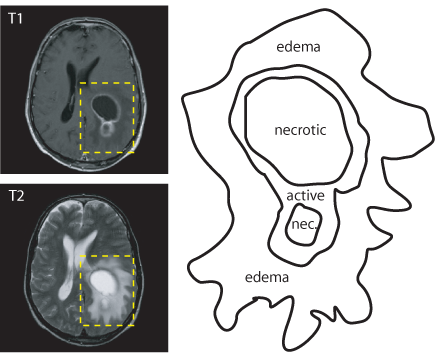 We focus specifically on glioblastoma multiforme brain cancer (GBM). For
example, over 40,000 patients are diagnosed annually with primary brain tumors
and approximately 40% of these are GBMs [Smirniotopoulos, 1999]; the median
postoperative survival time is extremely short (8 months) with a 5-year
recurrence- free survival rate of nearly zero [Patel and Tse, 2004]. However,
like most segmentation problems, automatic quantification of tumors is very
difficult. In general, it is impossible to segment a tumor by simple
thresholding techniques [Prastawa et al., 2004] because, although medical
images are constrained signals, the organic structures therein present many
modeling challenges [Patel and Tse, 2004]. The organic structures' geometry is
deformable, has high variation, and is often articulated. The appearance is
similarly complex, especially for pathological structures, due to the general
coarse resolution at which the images actual sample the underlying physical
processes.
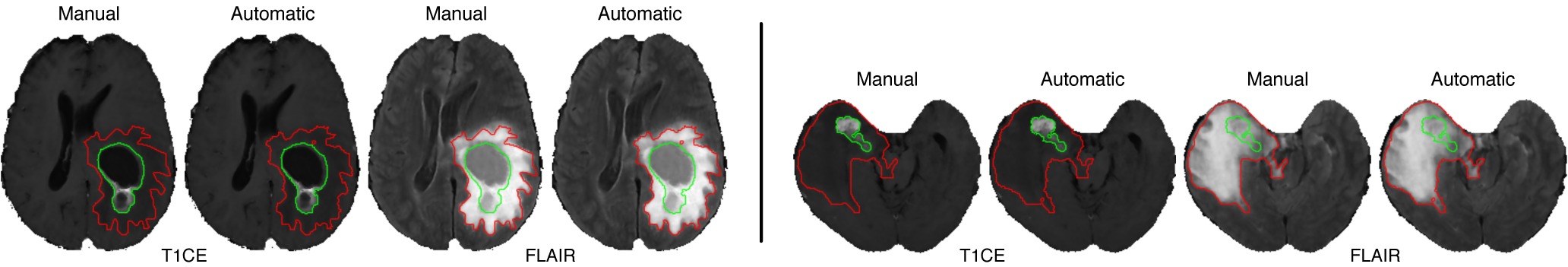
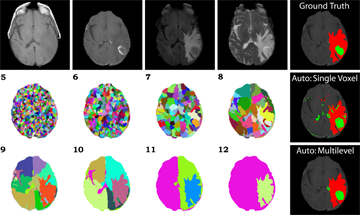
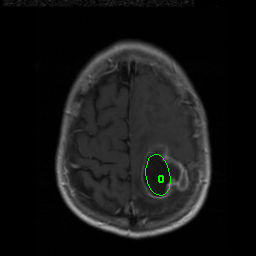
We have achieved very promising results using a combination of our multilevel segmentation with integrated Bayesian model classification, dicriminative machine learning techniques, and graph-shifts based energy minimization. Some examples are below.
We focus specifically on glioblastoma multiforme brain cancer (GBM). For
example, over 40,000 patients are diagnosed annually with primary brain tumors
and approximately 40% of these are GBMs [Smirniotopoulos, 1999]; the median
postoperative survival time is extremely short (8 months) with a 5-year
recurrence- free survival rate of nearly zero [Patel and Tse, 2004]. However,
like most segmentation problems, automatic quantification of tumors is very
difficult. In general, it is impossible to segment a tumor by simple
thresholding techniques [Prastawa et al., 2004] because, although medical
images are constrained signals, the organic structures therein present many
modeling challenges [Patel and Tse, 2004]. The organic structures' geometry is
deformable, has high variation, and is often articulated. The appearance is
similarly complex, especially for pathological structures, due to the general
coarse resolution at which the images actual sample the underlying physical
processes.
We have achieved very promising results using a combination of our multilevel segmentation with integrated Bayesian model classification, dicriminative machine learning techniques, and graph-shifts based energy minimization. Some examples are below.
 Other Information:
Publications
Other Information:
Publications
Past Members Alan Yuille (UCLA), Usha Sinha (UCLA), Shishir Dube (UCLA), Timothy Burns There is thus currently a great deal of interest in the establishment of imaging as a valid biomarker in the assessment of the response of cancer to a variety of treatments [Bradley et al., 2002, Tuma, 2006]. Of the possible measure- ments that can be made on images to assess the change in tumor burden, overall tumor volume is the key indicator [Bradley et al., 2002, Liu et al., 2005]. However, rather than using the full tumor volume, the current international standard measurements rely on basic quantitative surrogates such as the two dimensional World Health Organiza- tion (WHO) [World Health Organization, 1979] or Response Assessment Criteria for Solid Tumors (RECIST) one dimensional criterion [Therasse et al., 2000] used to measure patient response. Specifically, tumors are measured in one or two dimensions at their baseline�the area of maximal size, which is selected manually by the physician. These current quantitative techniques, although offering numerical representations of the state of the patient, are gross approximations to the true underlying tumor and are computed manually, hence time consuming and often not reproducible. They are therefore considered by many to be inadequate to allow imaging to serve as a biomarker on which to base treatment [Patz Jr. et al., 2000, Tuma, 2006]; they have generally failed to provide repeatable and statistically meaningful prognostic parameters of outcome prediction in both the clinic and the research lab. Hence, advanced image segmentation and recognition techniques are needed to improved the measurement science of cancer imaging and tumor staging.
 We focus specifically on glioblastoma multiforme brain cancer (GBM). For
example, over 40,000 patients are diagnosed annually with primary brain tumors
and approximately 40% of these are GBMs [Smirniotopoulos, 1999]; the median
postoperative survival time is extremely short (8 months) with a 5-year
recurrence- free survival rate of nearly zero [Patel and Tse, 2004]. However,
like most segmentation problems, automatic quantification of tumors is very
difficult. In general, it is impossible to segment a tumor by simple
thresholding techniques [Prastawa et al., 2004] because, although medical
images are constrained signals, the organic structures therein present many
modeling challenges [Patel and Tse, 2004]. The organic structures' geometry is
deformable, has high variation, and is often articulated. The appearance is
similarly complex, especially for pathological structures, due to the general
coarse resolution at which the images actual sample the underlying physical
processes.
We have achieved very promising results using a combination of our multilevel segmentation with integrated Bayesian model classification, dicriminative machine learning techniques, and graph-shifts based energy minimization. Some examples are below.
We focus specifically on glioblastoma multiforme brain cancer (GBM). For
example, over 40,000 patients are diagnosed annually with primary brain tumors
and approximately 40% of these are GBMs [Smirniotopoulos, 1999]; the median
postoperative survival time is extremely short (8 months) with a 5-year
recurrence- free survival rate of nearly zero [Patel and Tse, 2004]. However,
like most segmentation problems, automatic quantification of tumors is very
difficult. In general, it is impossible to segment a tumor by simple
thresholding techniques [Prastawa et al., 2004] because, although medical
images are constrained signals, the organic structures therein present many
modeling challenges [Patel and Tse, 2004]. The organic structures' geometry is
deformable, has high variation, and is often articulated. The appearance is
similarly complex, especially for pathological structures, due to the general
coarse resolution at which the images actual sample the underlying physical
processes.
We have achieved very promising results using a combination of our multilevel segmentation with integrated Bayesian model classification, dicriminative machine learning techniques, and graph-shifts based energy minimization. Some examples are below.



| [1] | W. Wu, A. Y. C. Chen, L. Zhao, and J. J. Corso. Brain tumor detection and segmentation in a CRF (conditional random fields) framework with pixel-pairwise affinity and superpixel-level features. International Journal of Computer Aided Radiology and Surgery, 9(2):241-253, 2014. [ bib | http ] |
| [2] | L. Zhao, W. Wu, and J. J. Corso. Semi-automatic brain tumor segmentation by constrained MRFs using structural trajectories. In Proceedings of Medical Image Computing and Computer Aided Intervention, 2013. [ bib | .pdf ] |
| [3] | S. Dube, J. J. Corso, A. Yuille, T. F. Cloughesy, S. El-Saden, and U. Sinha. Hierarchical Segmentation of Malignant Gliomas Via Integrated Contextual Filter Response. In Proceedings of SPIE Conference on Medical Imaging, 2008. [ bib ] |
| [4] | J. J. Corso, E. Sharon, S. Dube, S. El-Saden, U. Sinha, and A. Yuille. Efficient Multilevel Brain Tumor Segmentation with Integrated Bayesian Model Classification. IEEE Transactions on Medical Imaging, 27(5):629-640, 2008. [ bib | .pdf ] |
| [5] | S. Dube, J. J. Corso, T. F. Cloughesy, S. El-Saden, A. Yuille, and U. Sinha. Data Mining Systems Analysis and Optimization in Biomedicine, chapter Automated MR Image Processing and Analysis of Malignant Brain Tumors: Enabling Technology for Data Mining. American Institute of Physics, 2007. [ bib ] |
| [6] | J. J. Corso, A. L. Yuille, N. L. Sicotte, and A. Toga. Detection and Segmentation of Pathological Structures by the Extended Graph-Shifts Algorithm. In Proceedings of Medical Image Computing and Computer Aided Intervention (MICCAI), 2007. [ bib | .pdf ] |
| [7] | J. J. Corso, E. Sharon, and A. L. Yuille. Multilevel Segmentation and Integrated Bayesian Model Classification with an Application to Brain Tumor Segmentation. In Proceedings of Medical Image Computing and Computer Aided Intervention (MICCAI), volume 2, pages 790-798, 2006. [ bib | .pdf ] |